I am a scientist working at an antibody discovery company, where I apply machine learning and AI approaches to accelerate in silico antibody discovery and protein engineering.
My work focuses on leveraging protein language models, structure prediction tools, and ML pipelines to identify and engineer therapeutic antibody candidates.
This site showcases my research projects, technical skills, and thoughts on applied ML in biologics discovery.
I believe in the power of collaboration. If you see an opportunity for us to work together or need support, please reach out.
- Generative AI for Antibody/Protein Design
- Protein Structure Prediction
- Machine Learning for Therapeutic Discovery
- Multi-omics Data Integration
- Bioinformatics & NGS Analysis
Bachelor's in Neuroscience, Data Science and Psychology
Concordia College
Technical Skills
Experience
Lead in silico workflows for antibody discovery and characterization. Own end-to-end sequence-function modeling workflows.
diffusion models structure-function modeling antibody engineering PyTorch HPC
I build diffusion-based pipelines for de novo antibody sequence design and custom ML tools for predicting affinity, stability, and manufacturability from proprietary wet lab data. The work is about turning model outputs into testable hypotheses for the bench team.
diffusion models structure-function modeling antibody engineering PyTorch HPC
Built a deep learning humanization platform for optimizing mouse-derived lead candidates and led the onboarding of Oxford Nanopore sequencing infrastructure, including the downstream analysis workflows.
antibody humanization deep learning PLM NGS pipelines
Built data pipelines from production databases to identify manufacturing bottlenecks and reduce turnaround times for gene therapy products. Got a thorough look at GMP environments and the quality standards that govern therapeutics.
data pipelines gene therapy GMP manufacturing
Transcriptomic analysis on Nanopore RNA-seq data and image analysis pipelines in ImageJ for characterizing microglial morphology. Contributed to work published in Cell Reports Medicine.
RNA-seq analysis Nanopore Sequencing cloud compute ImageJ neuroimmunology
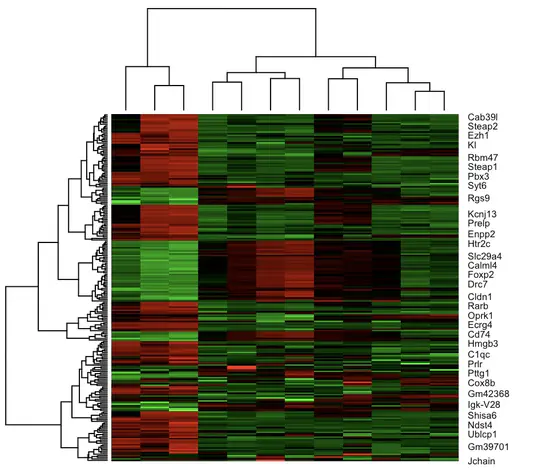
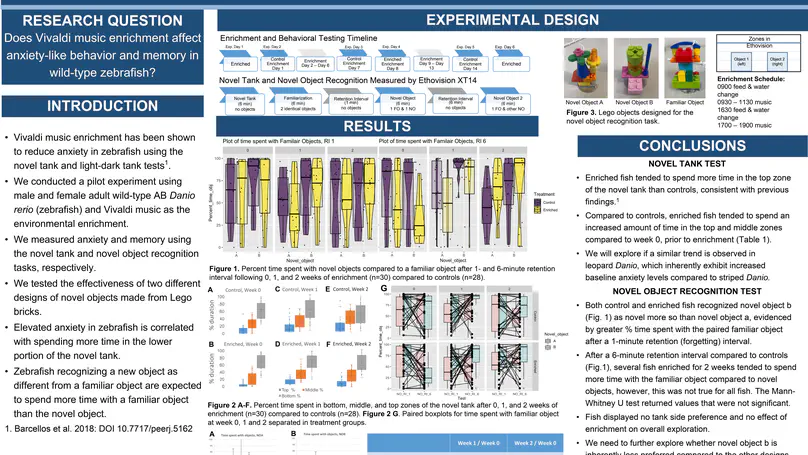
Designed zebrafish behavioral enrichment experiments, performed gene enrichment analysis in R, and presented at multiple academic conferences.
behavioral neuroscience R gene enrichment zebrafish anatomy
Certificates
Recent Posts
Projects
Contact
Feel free to contact me via email or LinkedIn.